THE AFFECTED GENE
FIBROBLAST GROWTH FACTORS
|
Fibroblast growth factors (FGF) are a family of growth factors that are involved in a variety of signalling pathways
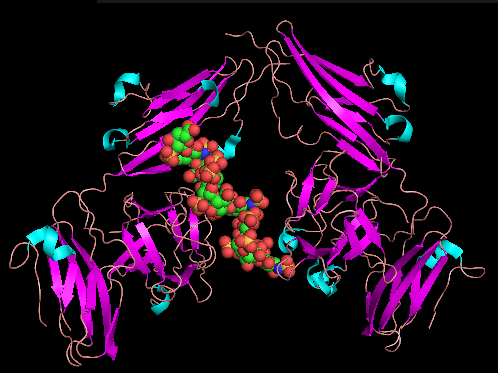
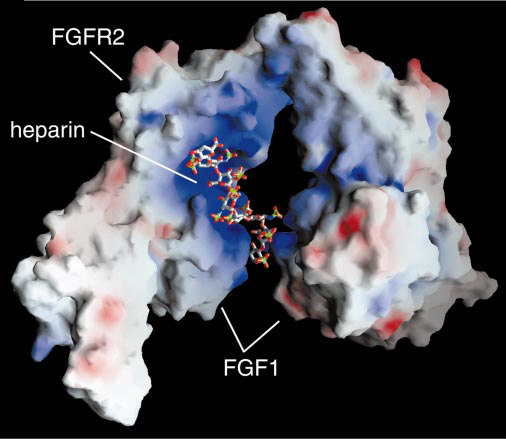
Fibroblast Growth factors are encoded by 22 genes. The molecular weight is 92,025 Da. [10] They have the potential to bind to four high affinity ligand dependent FGFR tyrosine kinase molecules (FGFR1-4), but only in the presence of herapin sulfate (HS) glycosaminoglycans. FGF's stably bind FGFRs and lead to the formation of 2:2:2FGF-FGFR-HS dimers which enable cytoplasmic kinase domains to transphosphorylate and become activated. [14] FGF signalling activity is critical for maintaining the balance between cell proliferation and differentiation, ensuring that no increased activity in FGFR2 activity occurs that could result in Apert Syndrome. Receptor binding specificity is an essential mechanism in the regulation of FGF responses and is primarily achieved through alternative splicing of 2 exons, IIB and IIC, in the second half of (D3) in FGFR. [5]
In terms of Evolutionary Conservation ,the most related orthologous and homologues of the FGFR2 gene and FGFR2 protein are in; Tupaia Chinesis, Bos Taurus, Oryctolagus Cuniculus, Gallus Gallus. Columba Livia, Xenopus Laevis and Danio Derio [11] |
|
THE MUTATION
|
Although the
effects of the AS missense mutations on receptor structure is not known, it is
likely that the AS mutations alter the conformation of the linker region and
change the relative orientation of Ig-like domains II and III. These mutations affect ligand binding through direct side chain
interactions. [17]
Most mutations in the extracellular domain of FGFR2 cause receptor activation by facilitating dimerization. AS is caused by substitution of one of 2 adjacent residues, Ser252Trp or Pro325Arg, in the highly conserved linker region between Ig-like domain 2 (D2) and (D3). AS mutations create additional interactions between FGFR and FGF that cause a conformational change increasing ligand-binding affinity, and decrease the dissociation rate of FGFR2 from FGF in vitro and cause ligand dependent FGFR2 activation in vivo.[18-19] The basis for increased affinity of FGF2 for the Ser252Trp mutant FGFR2(IIc) is the hydrophobic contact between Phe-21 of FGF2 and Trp-252, Tyr-281 and Ile-257 of FGFR2(IIc). [11] Therefore the Ser252Trp mutant S252W mutation in FGFR2 allows FGFR2c to be activated by FGF7 and FGF10. And for FGFR2b to be activated by FGF2,6,9 allowing autocrine signalling tissues that express these ligands. Mutations introduce additional interactions between FGFR2 and FGF2, thereby by augmenting FGFR2-FGF2 affinity It has been noted that
|